Story
Synthetic circuits for cell ratio control
Key takeaway
Scientists have created synthetic genetic circuits that can precisely control the ratios of different cell types, which could enable new methods for tissue engineering and regenerative medicine.
Quick Explainer
The researchers developed a suite of genetic devices that use recombinase enzymes to precisely control the ratios of multiple cell types derived from a single founder. These devices contain recombinase recognition sites that can be selectively excised, generating distinct cell populations expressing different reporters. By combining multiple branching devices, the researchers were able to create complex multicellular consortia with programmable differentiation patterns, enabling the bottom-up construction of synthetic multicellular systems. This modular, recombinase-based approach provides a versatile platform for applications ranging from metabolic engineering to tissue engineering, leveraging the ability to achieve user-defined, autonomous cell-type ratios.
Deep Dive
Technical Deep Dive
Overview
This work introduces a suite of genetic devices using recombinase enzymes to precisely control the ratios of multiple cell types derived from a single founder. The devices enable user-defined, autonomous differentiation programs, facilitating the bottom-up construction of synthetic multicellular consortia and coordinated multicellular assemblies.
Problem & Context
Recent advances in genetic engineering have enabled artificial diversification of cell populations. However, achieving precise control over the ratios of multiple cell types remains a major challenge. Natural multicellular systems accomplish sophisticated tasks through the differentiation and division of labor of specialized cell types.
Methodology
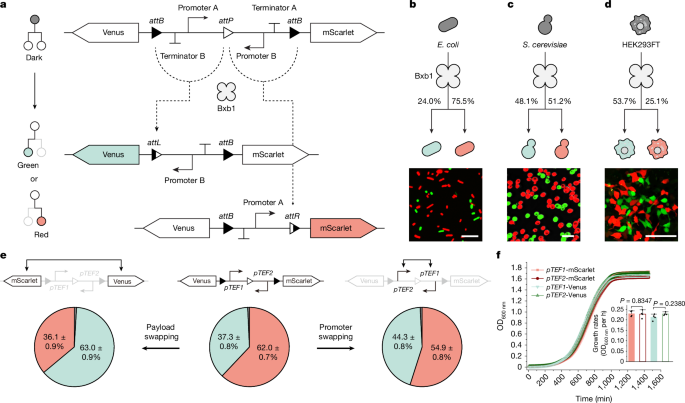
The researchers leveraged serine recombinase enzymes, such as Bxb1, to engineer binary cell-fate branching devices. These devices contained a single recombinase recognition site (attP) flanked by two identical attB sites. Recombinase-driven excision of one of the attB sites generated two distinct cell populations expressing different reporters.
Key parameters explored:
- Promoter sequences between att sites
- Length of DNA sequences flanked by att sites
- Variants of recombinase att sites
The researchers also combined multiple branching devices in parallel or series to generate complex multicellular populations.
Data & Experimental Setup
Experiments were conducted in representative bacterial (E. coli), yeast (Saccharomyces cerevisiae), and mammalian (HEK293FT, CHO-K1) systems. Device performance was characterized using flow cytometry, microscopy, growth assays, and computational modeling.
Results
- Precise control over binary cell-fate branching and population composition
- Ability to combine multiple branching devices to generate up to 8 distinct cell types
- Implementation of exponential functions to achieve extremely low ratios of specific cell types
- Engineering of sequential differentiation programs and self-organizing multicellular architectures
- Application to diverse consortia for pigment production, cellulose degradation, and morphogenesis
Interpretation
The recombinase-based branching devices provide a modular, expandable platform for programmable ratio-specific cell differentiation and pattern formation. The generality of the approach across species and its independence from pre-established morphogen or quorum signals make it a versatile tool for applications ranging from metabolic engineering to tissue engineering.
Limitations & Uncertainties
- Incomplete recombination leaves a reserved subpopulation of undifferentiated or partially differentiated progenitors
- Potential toxicity or differential growth rates of some recombinases
- Ability to maintain desired cell ratios over prolonged culture periods
What Comes Next
Future work may explore feedback mechanisms, quorum sensing, and less toxic recombinases to boost the robustness of the devices. Integrating additional signaling modalities could enable more complex, context-dependent differentiation programs for engineered organoids, tissues, and metabolic systems.